Introduction
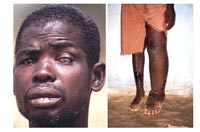
Filarial infections such as onchocerciasis and lymphatic filariasis are caused by tissue-invading parasitic worms of the phylum Nematoda. These infections lead to severe morbidity and considerable socio-economic problems throughout the tropics (Fig. 1). Filariases in total are still the third most important tropical disease worldwide according to their Disability Adjusted Life Years (DALY) as a measure of disease burden. Onchocerciasis is the second most common infectious blinding disease (river blindness) of the developing world afflicting 18 million people, 95% of whom live in West and Central Africa.
Epidemiologic evidence suggests a genetic component contributes to susceptibility and possibly the outcomes of filarial infection.
Animal models have a tremendous potential to dissect complex interactions of multiple genes that control the host response to a pathogen. Approaches to unravel these complexities include quantitative trait loci (QTL) mapping and positional cloning of host-resistance genes in mutant or congenic mice, an approach used widely for microbial infections but rarely for helminths.
In our project murine infections with the filarial nematode Litomosoides sigmodontis are used to study innate and adaptive immune responses to nematode infections and as a paradigm model for helminth-associated immunomodulation. L. sigmodontis is the only filaria known to undergo complete development in some inbred strains of mice.
BALB/c mice are susceptible while mice of the C57BL/6 or DBA/1 strain are resistant. In patently infected BALB/c mice, and as seen in human filarial infections, a Th2-type cytokine response predominates, initiating pathology as well as self-limiting the parasite load.
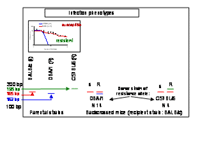
Different strains of mice were found to differ also widely in their capability to eliminate circulating microfilariae (1st larval stage of the worm; Fig. 2). The most extreme phenotypes showed DBA/1 mice being completely microfilariae-negative 2-3 days after injection of worm larvae and therefore assigned as resistant, and the BALB/c mice that tolerate the blood-circulating microfilariae for several weeks.
The F1 generation of DBA/1 as the donor strain for resistance and of susceptible BALB/c mice exhibited a uniform resistant phenotype. Backcross with BALB/c resulted in a division of 50% resistant versus susceptible animals. This trait of inheritance suggests that resistance to injected microfilariae is inherited dominantly by one gene or closely linked genes. In each generation from N2 to N14 the offspring was infected, then mf blood-levels were checked at day 4 p. i. and resistant animals were selected for further backcross with BALB/c.
Using such a serial backcross strategy followed by an intercross of N15 mice we finally obtained congenic mouse strains sharing the BALB/c genetic background but which differ profoundly in their susceptibilty to intravenously injected microfilariae.
Genes and gene products that define such differences were characterized by a dual approach. 1. Scanning of the genomes of backcrossed mice in different generations, of congenic mice and of the original parental strains by use of microsatellite analyses (in close collaboration with Dr. A. Lengeling, GBF Braunschweig, and Prof. P. Nuernberger, Dr. B. Meyer, Max-Delbrueck-Center, Berlin). 2. Gene expression using microarray analyses with high density DNA-chips was compared between resistant and permissive congenic mouse strains before and after infection.
Results
Genome-wide scanning of congenic mice by microsatellite analysis revealed a candidate region for the postulated microfilaraemia resistance locus on chromosome 12.
A second backcross approach was started with C57BL/6 mice as donor strain for resistance and BALB/c as recipient strain again. So far (N4 generation) a similar trait of inheritance was found with these parental strains. Moreover, preliminary results of microsatellite analysis revealed the same candidate region for microfilaraemia resistance in the C57BL/6 mice as in the DBA/1 strain. By use of two microsatellite marker we managed to prospectively genotype the resistant/ susceptible phenotype of backcrossed mice (Fig. 3).
Fine mapping of the candidate region and identification of the microfilaraemia resistance gene will be achieved by SNP analyses and positional cloning.
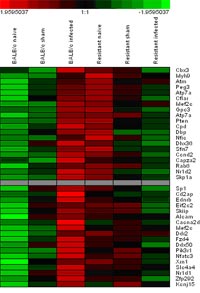
By determining gene expression profiles with high-density DNA microarrays (Affymetrix) we identified distinct set of genes differentially expressed between congenic mice (Fig. 4). Most genes were associated with host immune cell survival, apoptosis or proliferation or encode factors of adhesion or migration of host phagocytes. The biological significance of selected candidate genes was further characterized in vitro and in vivo by Real Time-PCR, immunological assays and use of appropriate gene knockout mouse models.
In order to further investigate the specificity of this innate immune response to microfilariae in our congenic mice we injected worm larvae of the human pathogenic species Brugia malayi and of the rodent filaria Acanthocheilonema viteae. These microfilariae differ profoundly in their surface structure (sheated versus unsheathed) as well as in the presence or absence of bacterial endosymbionts of the genus Wolbachia.
Interestingly, the same disparate outcome following infection with L. sigmodontis in our congenic mice could also be observed after injection of microfilariae of Brugia malayi but not with those of Acanthocheilonema viteae. These similarities in the innate immunity to microfilariae of Litomosoides and Brugia probably reflect the biochemical resemblances of their surface structures. No difference between the congenic strains could be observed after infection with the gastrointestinal nematode Angiostrongylus costa-ricensis, with the trematode Schistosoma mansoni, with the protozoan Plasmodium berghei ANKA or with the gram-negative Enterobacterium Yersinia enterocolitica (collaboration with Prof. I. Autenrieth and Dr. E. Bohn, Medical Microbiology, Tuebingen).
Another striking feature of helminth infections is the profound down-modulation of inflammatory reactions to both specific and non-specific stimuli 4.This pheno-menon was used in an approach to modify the allergic sensitization to Ovalbumin in a murine model for allergic asthma and airway hyperreactivity. In collaboration with Prof. E. Hamelmann and Dr. A. Dittrich, Pedriatic Pneu-mology & Immunology, Charité, Berlin, it could be de-monstrated for the first time that filarial infection suppressed all aspects of the asthmatic phenotype.
Outlook
Different inbred strains may be regarded as a model for different allelic variants of immune reponse genes in a human population. Therefore the identification of resistance/susceptibility gene in murine filarial infection could facilitate the search for homologous genes in human filariasis. The identification of susceptibilty genes in filarial infection could provide new insights into therapeutic strategies, including pharmacological intervention and vaccine development, and influence public health measures to control or avert infection.
Lit.: 1. Choi EH et al. (2003) Genetic variation in immune function and susceptibility to human filariasis. Expert Rev Mol Diagn 3: 367-74. 2. Lengeling A et al. (2001) The battle of two genomes: genetics of bacterial host/patho-gen interactions in mice. Mamm Genome 12: 262-71. 3.Hoffmann WH et al. (2000) Litomosoides sigmodontis in mice: reappraisal of an old model for filarial research. Parasitology Research 16:387-389.4. Hoffmann WH et al. (2001) Determinants for resistance and susceptibility to microfilaraemia in Litomosoides sigmodontis filariasis. Parasitology 122: 641-49.5. Dittrich AM et al. (2005) Infection with Litomosoides sigmodontis suppresses allergen-induced sensitization and pulmonary inflammation in a murine asthma model . Journal of Allergy & Clinical Immunology 115: 205.